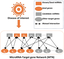
**Motivation:**
The misregulation of microRNA (miRNA) has been shown to cause diseases. Recently, we have proposed a computational method based on a random walk framework on a miRNA-target gene network to predict disease-associated miRNAs. The prediction performance of our method is better than that of some existing state-of-the-art network- and machine learning-based methods since it exploits the mutual regulation between miRNAs and their target genes in the miRNA-target gene networks.
**Results:**
To facilitate the use of this method, we have developed a Cytoscape app, named RWRMTN, to predict disease-associated miRNAs. RWRMTN can work on any miRNA-target gene network. Highly ranked miRNAs are supported with evidence from the literature. They then can be also visualized based on the rankings and in relationships with the query disease and their target genes. In addition, automation functions are also integrated, which allow RWRMTN to be used workflows from external environments. RWRMTN is among the first GUI-based tools for the prediction of disease-associated miRNAs.